 Star products
Star products
 Follow Us
Follow Us

Scan and follow us
Learn more
| Tool | Function introduction | Provide products | |
|---|---|---|---|
| mKeima | Study on mitophagy phenotype |
LV/Ad/AAV | |
| Cox8-EGFP-mCherry |
|||
| mito-dsred |
EGFP-LC3 |
||
| Parkin-EGFP |
Study on mitophagy pathway | ||
| Bnip3-EGFP |
|||
| Nix-EGFP |
|||
| FUNDC1-EGFP |
|||
1、 Pink/Parkin pathway
2、 Bnip3/Nix pathway
3、 FUNDC1 pathway
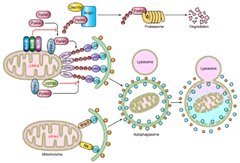
Mitophagy pathway: Pink1/Parkin OR Bnip3/Nix
Abnormalities in mitophagy are closely related to many diseases, so it is biologically important to study the specific molecular mechanisms of mitophagy as well as its physiological significance. HANBIO has an experienced mitophagy research team to provide you with a research program and tools for mitophagy.
1)Ad-GFP-LC3 adenovirus system, which can efficiently infect target cells to express GFP-LC3, and the overall level of autophagy can be observed in real time under fluorescence microscope in the infected cells;;
2)Ad-Dsred-Mito adenovirus system, a mitochondria-specific localization fluorescent probe (HBAD-Mito-dsred) developed independently by HANBIO can accurately localize and label mitochondria, which can be used in combination with HANBIO's exclusive dual-fluorescent LC3 cellular autophagy adenovirus, it can accurately track the dynamic process of mitophagy in real time;
Usage: Ad-GFP-LC3 + HBAD-Mito-dsred co-infected target cells, confocal detection of dual fluorescence co-localization, and the presence of mitophagy if co-localized!
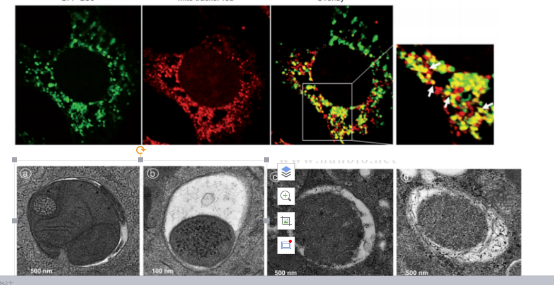
3)mKeima virus tools: The Keima protein expressed by mKeima is localized in the mitochondrial matrix, and when the mitochondrial autophagosome fuses with acidic lysosomes, the fluorescence signal of Keima protein shifts from green to red, and the conversion of the fluorescence signal can quantitatively reflect the occurrence and development of mitophagy.

PMID: 33042281
4)Cox8-EGFP-mCherry virus tools: The leading peptide sequence of the mitochondrial inner membrane protein Cox8 was expressed in fusion with the EGFP-mCherry fluorescent tandem sequence, and in the cytoplasm, both red fluorescent proteins and green fluorescent proteins were simultaneously expressed, appears as yellow. If mitophagy occurs, the green fluorescent protein is quenched in autophagic lysosomes due to the acidic environment of lysosomes, leaving only the red fluorescent signal. The level of mitophagy can therefore be measured by counting the number of red fluorescent spots.
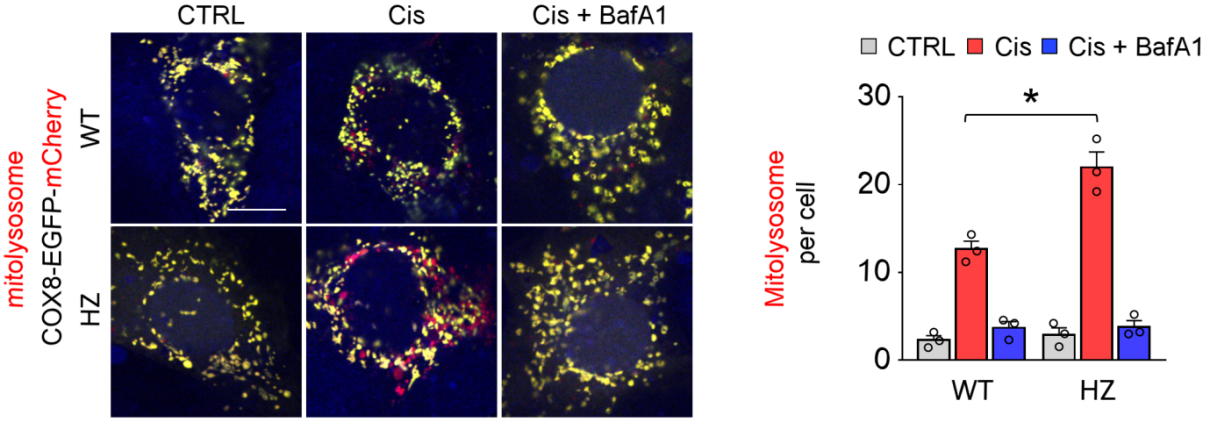
PMID:34739325
HANBIO mitophagy pathway research tool
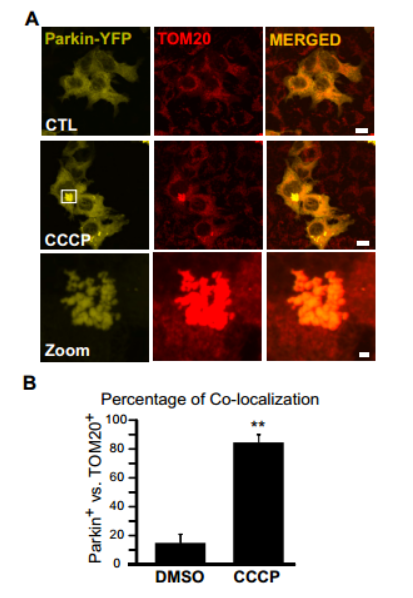
1)Ad-Parkin-EGFP
2)Ad-Bnip3-EGFP+Ad-Nix-EGFP
3)Ad-FUNDC1-EGFP
Usage: With the HANBIO HBAD-Mito-dsred to localize mitochondria co-infecting target cells, confocal detection of co-localization, and identification of mitochondrial translocation of relevant signaling molecules!
In addition, we also have adenovirus spot Beclin1, Atg7 and other common genes involved in autophagy to meet the needs of your experiment.

联系我们

返回顶部